MAP-Xing the malaria protein complexes
An international research team headed by the Bozdech laboratory at the Nanyang Technical University in Singapore and the Gilberger laboratory at the Centre for Structural Systems Biology and Bernhard-Nocht Institute for Tropical Medicine in Germany reveals the global landscape of protein-protein interactions that govern the biology of the malaria parasite. Their new approach named meltome-assisted profiling of protein complexes (MAP-X) was recently published in in Nature Microbiology.
Resulting in over half a million deaths each year, malaria remains a major burden for global health. The deadliest form of malaria is caused by the parasite Plasmodium falciparum. Considerable efforts have been undertaken to develop and deploy anti-malarial drugs, leading to the decrease of case fatality. Unfortunately, these advances are threatened by the rise of drug-resistant parasites. The development of novel therapeutics requires a better understanding of the cellular and molecular biology of the parasite itself.
Plasmodium falciparum is composed of over 5,200 distinct proteins and relies on a dynamic network of protein–protein interactions throughout its complex lifecycle.
“To understand the molecular mechanisms driving the complex biology of the parasite, it’s important to understand not only the role played by each individual protein but also their interactions with each other,” says Tim Gilberger who heads the Cellular Parasitology Department at BNITM. “However, the function and molecular interactions of over 40 percent of Plasmodium falciparum’s proteins are currently unknown, not to mention the uncharted territory of their dynamic interactions and complex formation during parasite multiplication.”
To discover new protein interactions, the researchers utilised a state-of-the-art method known as thermal proteome profiling (TPP).

“TTP examines the thermal stability of proteins in intact cells,” explains Samuel Pazicky, the paper’s first author “When exposed to heat, proteins that interact with one another are destroyed in a similar manner."
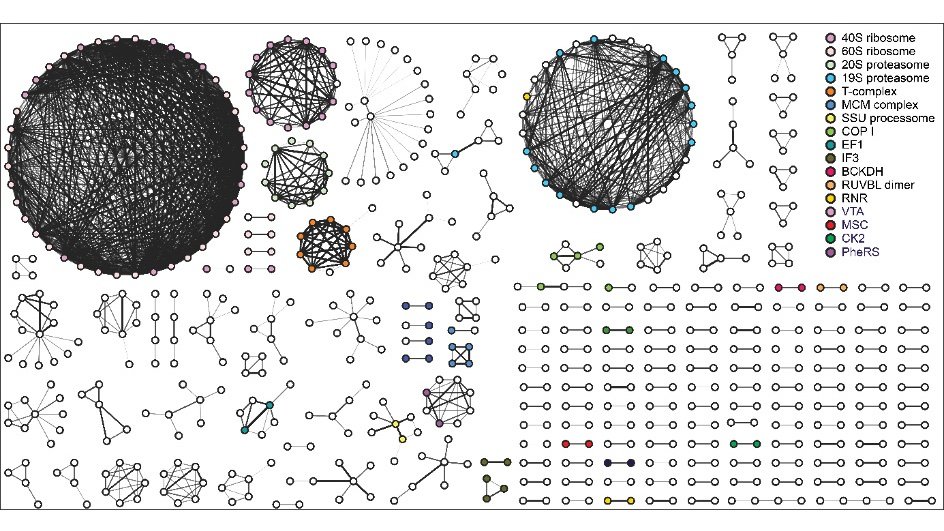
The research team then combined TPP with artificial intelligence to map and predict the protein-protein interactions. This new approach, called meltome-assisted profiling of protein complexes (MAP-X) applies machine learning to TPP data to monitor and compare thousands of proteins as they are expressed in cells. “We applied MAP-X across seven timepoints in the Plasmodium falciparum life cycle in human blood discovering more than 20,000 interactions,” says Pazicky.
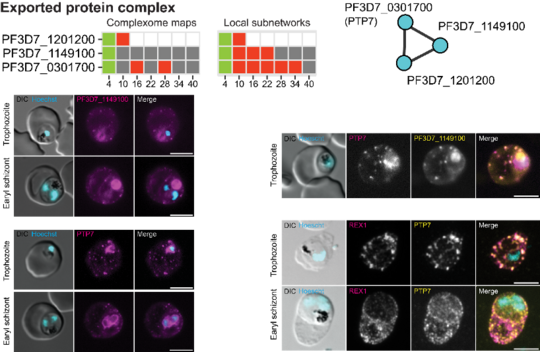
With MAP-X, the team not only confirmed the existence of known protein complexes but also discovered blueprints for novel parasite specific protein complexes and biochemical pathways. “The interaction maps generated by MAP-X were able to predict previously unknown protein complexes,” says Zbynek Bozdech, the papers corresponding author. “The existence of several of these complexes was further verified experimentally via fluorescence microscopy and pull-down assays.”
“With its ability to identifying previously undescribed interactions as well as reveal stage-specific dynamics, MAP-X is powerful resource for deciphering the dynamic interactions and fundamental biological processes of the malaria parasite,” explains Gilberger. MAP-X is expected to help guide the design of next-generation antimalarials at a time when resistance is threatening existing treatments. Outside of malaria research, MAP‑X represents a new method for mapping of protein complexes in any type of cell, including bacteria, other parasites as well as blood, plants or tissues.
Original publication
Pazicky S. et al.: MAP-X reveals distinct protein complex dynamics across Plasmodium falciparum blood stages. Nat Microbiol. (28 Nov. 2025). doi.org/10.1038/s41564-025-02173-7
Contact person
Prof. Dr Tim-Wolf Gilberger
Cellular Parasitology Department
Phone : +49 40 8998 87600
Email : gilberger@bnitm.de
Melissa Prass
Public Relations Officer CSSB
Phone : +49 40 8998-87502
Email : melissa.prass@cssb-hamburg.de
Julia Rauner
Public Relations
Phone : +49 40 285380-264
Email : presse@bnitm.de
Further information






