New detection method uses blood donations as an early warning system for West Nile virus
Researchers at the Bernhard Nocht Institute for Tropical Medicine (BNITM) have developed a new sequencing method that can detect the West Nile virus in blood donations. The highly sensitive method makes it possible to detect even tiny amounts of the virus and analyse its full genetic sequence. This enables researchers to identify virus variants at an early stage and track their spread in Germany. The researchers also showed that the virus was introduced into Germany several times independently from Central Europe and is now established in several eastern federal states. The study was recently published in the Journal of Infection.
![[Translate to English:] Rekonstruktion der Ausbreitung des West-Nil-Virus in Deutschland [Translate to English:] Die Grafik zeigt eine Europakarte, auf der in roten Linien die Verbreitungswege des West Nil-Virus eingezeichnet sind. Grün markierte Bereiche zeigen Regionen, in denen das Virus besonders aktiv war.](/fileadmin/_processed_/3/a/csm_Map_neu_7728bb41d9.jpg)
Mosquitoes transmit the West Nile virus (WNV), which in rare cases can cause severe neurological diseases in humans. Since its discovery in Germany in 2018, the virus has spread across several eastern federal states. Accurate genetic monitoring (genome surveillance) is crucial for detecting chains of infection and virus variants.
"The earlier we detect the virus, the faster health authorities can respond and prevent outbreaks," says Dr Dániel Cadar, head of the Virus Metagenomics and Evolution working group at BNITM and last author of the publication. "Genomic surveillance is our early warning system. It also shows us when and where new virus variants are becoming established."
New method detects even tiny amounts of virus
Cadar's team, together with cooperation partners from other institutes, has developed a new, highly sensitive method for detecting West Nile virus genetic material. Using this amplicon-based sequencing approach, the researchers can reliably detect even tiny amounts of viral genetic material and analyse the complete viral genome. Previously, metagenomic methods were used for detection, which analyse all genetic material in a sample and detect all DNA and RNA fragments from viruses, bacteria and cells at the same time. However, this method reaches its limits when virus quantities are very low and can no longer reliably detect the virus.
The new method is based on targeted "amplification reactions" (amplicons) that selectively amplify specific parts of the viral genome. This enables scientists to successfully analyse samples with very low viral loads.
"Our method enables genome sequencing even when only a few virus particles are present in the sample. Until now, this has been virtually impossible. Compared to metagenomic methods, our method is up to a hundred times more sensitive," says Cadar. "This lays the foundation for fast, accurate and cost-effective virus monitoring."


Blood donations as an early warning system
The new sequencing method makes it possible for the first time to successfully evaluate blood donations containing very small amounts of the virus. For this reason, BNITM researchers worked closely with the German Red Cross on the study. Blood donation samples are particularly valuable because they are collected nationwide, regularly and independently of disease symptoms. This provides a unique picture of virus circulation in the population. First author Gábor Endre Tóth explains:
"Blood donations are ideal early warning systems. They give us an accurate picture of where and when the virus is circulating, often before clinical cases occur. They combine routine health care with epidemiological surveillance. Thanks to this data, we can trace chains of infection and understand the spread of the virus much better."
On the trail of the virus
The researchers genetically analysed 43 blood donations that tested positive for West Nile virus between 2020 and 2024. This was more than ever before in Germany. The researchers "tracked" the virus genetically, allowing them to determine where a virus originated, how it has changed and which variants are currently in circulation. The genetic analyses show that the West Nile virus was introduced into Germany several times independently from Central Europe. This means it was not a single event, but rather the virus arrived in Germany via different routes and at different times. The new method also revealed that six different virus subgroups ("subclades") of the WNV lineage 2 have emerged. These variants are now established in several regions of Germany, especially in Berlin, Brandenburg, Saxony and Saxony-Anhalt.
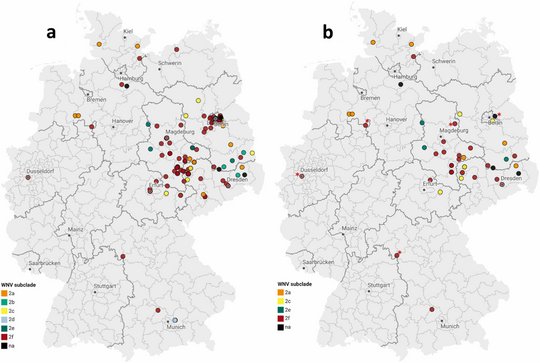
"Our study provides the most comprehensive overview to date of the genetic diversity and regional distribution of West Nile virus in Germany. The virus has long since become established here and has continued to evolve," said Tóth. "With our method, we can now track precisely and in real time how the virus adapts, how it migrates and which genetic lines prevail."
Foundation for future health monitoring
The study proves that the combination of sensitive sequencing technology and blood donation monitoring is an effective tool for the early detection, tracking and assessment of virus outbreaks. This applies not only to the West Nile virus, but also to other pathogens transmitted by mosquitoes.
"This approach significantly strengthens public health in Germany," says Cadar. "We can detect virus entries early, track their development and better understand their regional spread. This enables health authorities to respond more quickly and better prepare Germany for future outbreaks."
The method developed at BNITM and the associated bioinformatic pipeline are publicly accessible in the supplementary material of the manuscript. This will enable other laboratories in Europe to use the method in the future to detect virus outbreaks at an early stage. The researchers emphasise the importance of continuous and coordinated surveillance of mosquito-borne and other insect-borne pathogens, as climate change is likely to further increase their spread.
Original publication:
Tóth, G.E. et al., Blood donors as sentinels for genomic surveillance of West Nile virus in Germany using a sensitive amplicon-based sequencing approach, Journal of Infection 2025, DOI: 10.1016/j.jinf.2025.106647
Funding:
Federal Ministry of Research, Technology and Space (BMFTR)
Federal Ministry of Agriculture, Food and Regional Identity (BMLEH)
Contact person
Dr Dániel Cadar
Research Group Leader
Phone : +49 40 285380-840
Fax : +49 40 285380-400
Email : cadar@bnitm.de
Dr Anna Hein
Public Relations
Phone : +49 40 285380-269
Email : presse@bnitm.de
Further information






