Master Project Opportunities
Our research team offers a diverse range of master's project opportunities, spanning areas such as Next-Generation Sequencing (NGS) data analysis, evolutionary biology, transcriptomics, and artificial intelligence (AI). These projects involve tool development and/or infection biology research. If any of these areas spark your interest, please don't hesitate to contact us.
While each project requires specific expertise, we generally seek highly motivated students with strong coding abilities and a willingness to learn new skills within a collaborative team environment.
Project Descriptions:
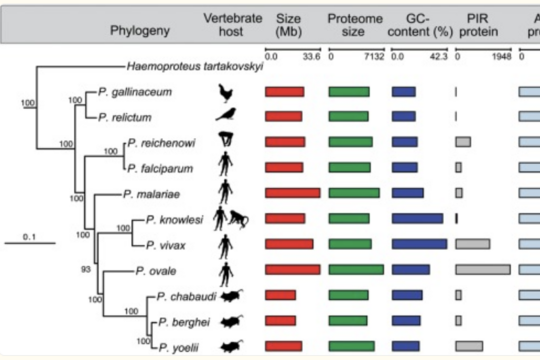
Analysis of a Novel Plasmodium Species
In collaboration with Franck Prugnolle and Laurent Reina, we have obtained sequencing data from a newly discovered rodent malaria species. This project aims to characterise this novel species, compare it to other rodent malaria species, and link observed phenotypes to specific genes. The project can begin with raw sequencing reads or utilise an assembled and annotated draft genome, depending on your interests.
Required Skills: Basic Linux command-line proficiency, understanding of evolutionary principles.
Transformer-Based Prediction of Gene Knockouts in Malaria
This project explores the application of transformer models, such as GeneFormer, to analyse malaria data. We have accumulated a substantial collection of single-cell transcriptomics datasets that can be used to train a transformer model for constructing gene regulatory networks. The goal is to determine whether our model can accurately predict the effects of gene knockouts.
- Required Skills: Proficiency in Python, a foundational understanding of AI principles (the more, the better), and a strong interest in parasitology.
Malaria niches and communication
The malaria parasite plasmodium affects tissue environments within their hosts to evade the immune system. At the same time, certain metabolic environments are linked to severity of infections. While immune system modulation is reported, the extent of host-parasite interaction is not well understood and spatial resolution of tissue environments in proximity of the parasite has long been missing to elucidate its details. By utilizing state-of-the-art spatial and single-cell transcriptomics we focus on the interaction between Plasmodium and its surrounding tissue niches, aiming to reveal crucial pathways during infection. A successful Master’s student will apply and improve analytic pipelines for spatial transcriptomics data of malaria infection, define molecular profiles of cells in infection niches, and propose important interaction pathways which could work as novel drug targets.
The ideal candidate’s profile:
- BSc in Bioinformatics, Molecular Life Sciences, Biology, Informatics or similar
- Experience in R or Python
- Interest in data and data visualization
- Experience in cell biology, immunology and/or disease mechanisms is a plus
- Motivation to work in an international team towards a common goal
- Good organisational and communication skills
- Creative mindset and strong problem-solving capabilities is a plus
- Experience in genetics NGS is a plus
- Ability to read, write, and communicate in English
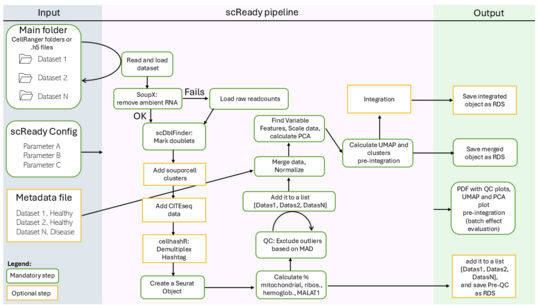
Exploring robust single cell pipelines
Systematic differences in sequencing coverage between libraries are often observed in sequencing data and arise from technical differences in how the data was achieved. Proper preprocessing is essential for many types of transcriptomic data and includes multiple steps that control for data quality, structure, and control for underlying biases, technical differences or batch effects. Various approaches have been developed for different types of transcriptomic data, which presents us with a problem if we aim to integrate datasets of different origin. Gene count-based methods introduce region- or cell type-specific biases that compromise downstream analyses in spatial data. Establishing reliable preprocessing pipelines is of highest interest within the scientific community and beyond. A successful Master's student will define the technical peculiarities of different transcriptomic techniques, describe the effects of different data processing methods onto integration, and develop solutions to minimise their impact on subsequent analyses.
The ideal candidate’s profile:
- BSc in Bioinformatics, Statistics, Data Science, Informatics or similar
- Competent in R or Python or similar coding language
- Interest in complex data and statistical problems
- Creative mindset and strong problem-solving capabilities
- Motivation to work in an international team towards a common goal
- Good organisational and communication skills
- Experience in cell biology, immunology and/or disease mechanisms is a plus
- Experience with transcriptomic data is a plus
- Ability to read, write, and communicate in English

Not sure, but this might give YOU a project. (c) risingscholars.net
Exploring robust single cell pipelines
Systematic differences in sequencing coverage between libraries are often observed in sequencing data and arise from technical differences in how the data was achieved. Proper preprocessing is essential for many types of transcriptomic data and includes multiple steps that control for data quality, structure, and control for underlying biases, technical differences or batch effects. Various approaches have been developed for different types of transcriptomic data, which presents us with a problem if we aim to integrate datasets of different origin. Gene count-based methods introduce region- or cell type-specific biases that compromise downstream analyses in spatial data. Establishing reliable preprocessing pipelines is of highest interest within the scientific community and beyond. A successful Master's student will define the technical peculiarities of different transcriptomic techniques, describe the effects of different data processing methods onto integration, and develop solutions to minimise their impact on subsequent analyses.
The ideal candidate’s profile:
- BSc in Bioinformatics, Statistics, Data Science, Informatics or similar
- Competent in R or Python or similar coding language
- Interest in complex data and statistical problems
- Creative mindset and strong problem-solving capabilities
- Motivation to work in an international team towards a common goal
- Good organisational and communication skills
- Experience in cell biology, immunology and/or disease mechanisms is a plus
- Experience with transcriptomic data is a plus
- Ability to read, write, and communicate in English
Integration of images with single cell
Cells undergo different states, depending on tissue environment, developmental state, cell-interaction, metabolism, function and lineage. For a long time cells were primary classified by morphology in order to study many questions in biology, immunology and disease. Today, RNA or surface marker profiling is used to classify cells and identify novel subclasses and cell states. With large amounts of imaging, sequencing, and other data of cell types in diverse states and under various conditions we aim to integrate different data types for a better understanding of cell states and a more fine grained classification of subtypes. A successful Master’s student will utilize modern machine learning techniques to classify cell morphologies in the context of infection and immune responses, integrate different data types and draw connection between cell morphologies and molecular cell states.
The ideal candidate’s profile:
- BSc in Bioinformatics, Data Science, Molecular Life Sciences, Biology, Informatics or similar
- Experience in R or Python or similar coding language
- Experience in machine learning is a plus
- Experience in cell biology, immunology and/or disease mechanisms is a plus
- Experience with transcriptomic data is a plus
- Creative mindset and strong problem-solving capabilities
- Motivation to work in an international team towards a common goal
- Good organisational and communication skills
- Ability to read, write, and communicate in English
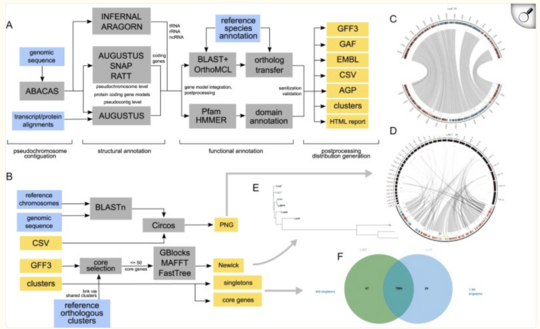
Add a function to the Companion Web Server
The Companion Webser is a well-established service used to annotate genomes of fungi, vectors, and parasites. It has a robust backend and is widely used, providing valuable outputs for researchers. However, to meet the evolving needs of the scientific community, several new functions could be implemented during this project. These additions would enhance the tool's capabilities, making it more versatile and powerful for genome analysis.
The ideal candidate’s profile:
- BSc in Bioinformatics, Data Science, Molecular Life Sciences, Biology, Informatics or similar
- Very strong coding background
- Deep understanding of software architecture
- Experience in de novo assembly and annotation is a plus
- Experience with transcriptomic data is a plus
- Creative mindset and strong problem-solving capabilities
- Motivation to work in an international team towards a common goal
- Good organisational and communication skills
- Ability to read, write, and communicate in English
Please contact us if you are interested. Also these is just a selection, and we are open for your own ideas.