Our research projects at a glance
We study the cell biology of the most virulent human malaria parasite, Plasmodium falciparum. By combining advanced experimental genetics with quantitative microscopy and chemical biology, we gain insights into the fundamental adaptations that safeguard the survival of these fascinating single cell organisms during blood stage development.
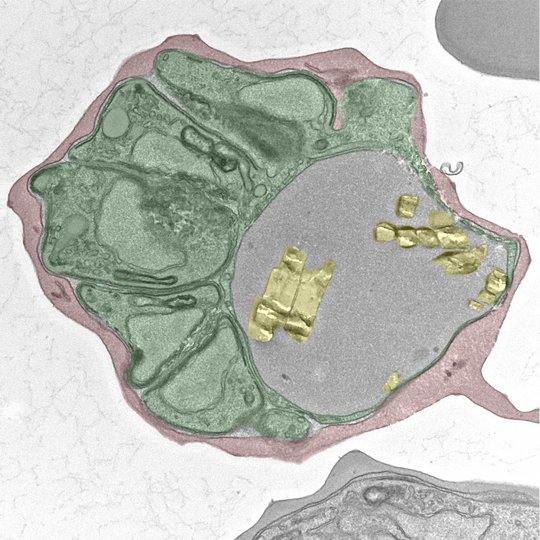
Malaria is a mosquito-borne disease that puts half of the world’s population at risk of infection. Despite rigorous control programs, malaria eradication remains one of the most testing medical challenges of the 21st century. Owing to the rapid emergence and spread of drug resistance, innovative intervention strategies are urgently needed and strictly depend on our improved understanding of the complex biology underlying this devastating disease. Malaria is caused by unicellular Plasmodium parasites (green), which repeatedly invade and lyse red blood cells in the human bloodstream. Throughout its development within the erythrocyte, the parasite endocytoses ~80% of the host cell cytoplasm (red), which is then catabolized in an acidic digestive vacuole (gray). Proteolytic cleavage of hemoglobin releases the co-factor heme, which is highly toxic in its unbound form. To avoid cell damage, the parasite sequesters the host-derived heme into bioinert crystals known as hemozoin (yellow), which accumulate in the digestive vacuole. The transition from hemoglobin to hemozoin is essential for parasite survival and is exploited in antimalarial chemotherapy. In our group, we study the physiology and molecular machinery of the parasite’s digestive vacuole as well as other parasite adaptations that are essential for blood stage survival. By combining Cas9-mediated genetic engineering with a variety of quantitative imaging approaches and chemical biology, we hope to gain a better understanding of the complex adaptations that ensure the survival of P. falciparum in the human bloodstream.
Hemozoin formation
The formation of hemozoin crystals is an essential and druggable parasite pathway. However, in the absence of parasite-centered research approaches, the mechanistic basis of this unique biomineralization process remains elusive.
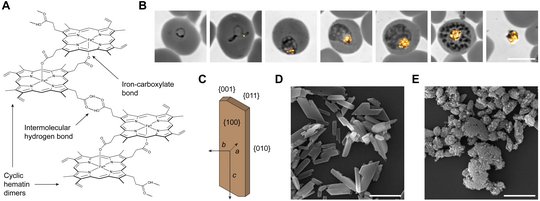
When the parasite proteolytically cleaves the hemoglobin inside the digestive vacuole, the iron-containing co-factor heme is released and rapidly oxidized to hematin. Owing to its pro-oxidant and detergent-like properties, free hematin is highly toxic. Thus, to avoid cell damage, two hematin monomers are linked via reciprocal iron-carboxylate bonds and the dimers assemble into high aspect ratio crystals through intermolecular hydrogen bonding (panel A). The formation of hemozoin is initiated during early parasite development and continues throughout parasite maturation. Upon parasite egress, the crystals are left behind, encapsulated in a residual body (panel B).
Although Robert Virchow and Charles Laveran described the connection between hemozoin and Plasmodium more than a century ago, the mechanisms of heme biomineralization remain elusive. Some studies suggest that hemozoin formation is a spontaneous and autocatalytic process, while others indicate an involvement of parasite proteins and lipids in the nucleation and growth of the crystals. The uncertainty surrounding this issue is alarming, considering that heme biomineralization is the target of the probably most effective antimalarial drug ever, chloroquine. This 4-aminoquinoline interferes with the adsorption of hematin onto growing hemozoin crystals, thereby causing a toxic build-up of unbound hematin. Although the global spread of efflux-mediated drug resistance has limited the utility of chloroquine, the outstanding efficacy of the 4-aminoquinoline antimalarials highlights hemozoin formation as a central Achilles’ heel of Plasmodium. If we can understand the underlying mechanisms, then there is hope for the development of similarly effective drugs targeting this unique parasite pathway. Therefore, we aim to identify and characterize the physicochemical and biogenic factors driving hemozoin formation in P. falciparum. Our studies prioritize experimentation with live malaria parasites over minimalistic crystallization assays and have previously led to the identification of a vacuolar lipocalin, called PV5/PfLCN, that modulates the quantities and architecture of hemozoin in cellulo (panels C-E). Elucidating the molecular functions of such parasite-tailored proteins will lead to a more complete understanding of this unique biomineralization process and might open up new avenues for antimalarial drug development.
Transport proteins of the digestive vacuole membrane
The digestive vacuole membrane of the malaria parasite is a critical interface that regulates vacuolar physiology and modulates antimalarial drug susceptibility.
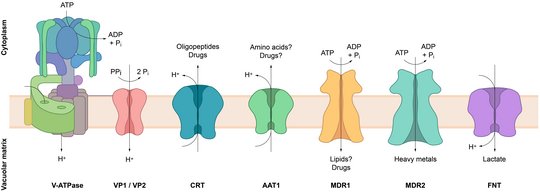
The membrane of the digestive vacuole harbors various transporters, which facilitate the exchange of ions, metabolites and (possibly) lipids. Some of these proteins, such as the chloroquine resistance transporter (CRT) and multi drug resistance protein 1 (MDR1), are important mediators of antimalarial drug resistance. The physiological function of CRT is the export of peptides derived from vacuolar hemoglobin catabolism, but resistance-confering mutations alter the transporter's substrate specificity and allow the efficient efflux of chloroquine, thereby removing the drug from its site of action. By contrast, MDR1 transports a wide range of substrates into the vacuolar matrix, including antimalarial drugs that act inside the digestive vacuole, such as chloroquine, and various drugs that act outside the vacuole. MDR1 modulates resistance towards these drugs through specific point mutations or through variation of its gene copy number. The rapid spread of resistance-conferring CRT and MDR1 mutations has led to a devastating efficacy loss of the former "wonder-drug" chloroquine and resulted in the global discontinuation of chloroquine monotherapy. Other vacuolar transporters, such as the V-type ATPase modulate the ionic milieu and pH of the digestive vacuole matrix, which is believed to affect vacuolar drug uptake.
Together, this underscores the digestive vacuole membrane as a critical interface that modulates the drug susceptibility of malaria parasites. The characterization of its functional molecules and transport pathways will enable us to understand and combat antimalarial drug resistance. Therefore, we characterize the physiological functions of known and novel transporters of the digestive vacuole membrane and study their roles in the transport of antimalarial chemotherapeutics.
Functional characterization of novel drug targets
Plasmodium-specific proteins that are essential for blood stage survival are attractive targets for therapeutic intervention. Their inhibition will only harm the parasite, not the host.
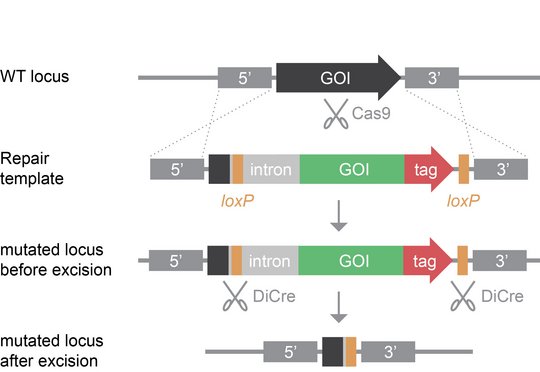
Nearly half of the parasite’s proteome comprises proteins of unknown function. Many of these are specific to the genus Plasmodium and do not contain functional annotations. The characterization of such enigmatic proteins, albeit challenging, offers intriguing new insights into the unique biology of Plasmodium parasites. Using state-of-the-art genetic systems, we study the localization and function of previously neglected Plasmodium-specific proteins with the goal to characterize them as potential new drug targets. Our genetic approach is predominantly based on Cas9- and DiCre-mediated genome editing. In the DiCre system, Cre recombinase is constitutively expressed as two enzymatically inactive domains, each fused to small rapamycin-binding proteins. Exogenously supplied rapamycin rapidly triggers hetero-dimerization of the DiCre proteins, reconstituting active Cre recombinase which can now excise chromosomal DNA sequences flanked by specific recombination motifs called loxP sites. Hence, introduction of two loxP sites into the gene of interest allows for rapid rapamycin-induced genetic ablation in a single intraerythrocytic cycle. Using this genetic system, the observation of rapamycin-induced parasite mortality currently provides the best evidence for gene essentiality, enabling the identification and functional characterization of novel drug targets.