Vision
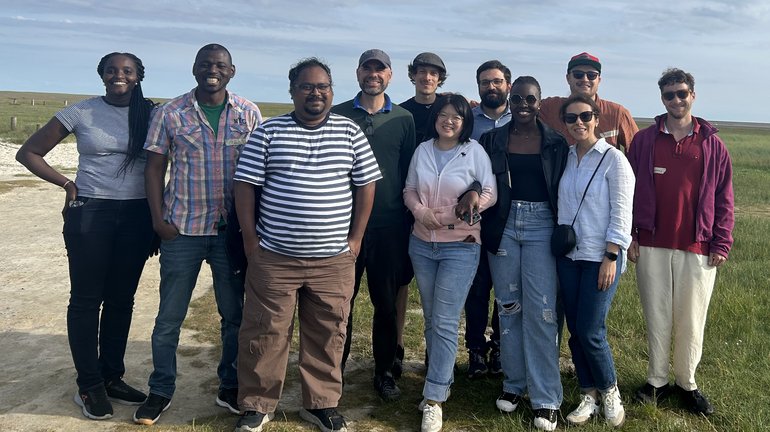
Our team, based in Hamburg and Glasgow, aims to advance understanding of host–pathogen interactions at the single-cell level to identify novel therapeutic targets.
Workstreams
- Pathogen evolution and immune evasion
We investigate the evolution of gene families that enable pathogens to evade or manipulate the host immune system. This includes generating and annotating genomes across diverse pathogens, as well as modelling complex gene families such as var genes in malaria. - Single-cell and spatial host–pathogen interactions
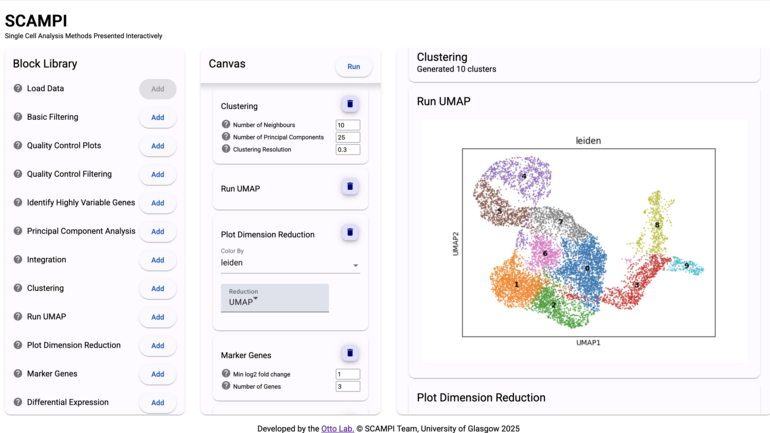
We integrate single-cell transcriptomics and spatial omics data to study how pathogens interact with host cells, and how the immune system responds. A particular focus is on understanding variability in immune responses across individuals. - Computational methods and tools
We develop and apply computational approaches—including machine learning and modern AI methods—to support these research areas and to empower our collaborators.
Data science support
In parallel, we actively support colleagues at BNITM (e.g. through drop-in sessions and monthly data Datathons - intranet) and collaborate globally through joint projects, training initiatives, and workshops, including the Hamburg Bioinformatics Summer School.
We place particular emphasis on partnerships in the Global South, aiming to strengthen the use of data science across diverse research contexts.